Learning from graph and relational data plays a major role in many applications including social network
analysis, marketing, e-commerce, information retrieval, knowledge modeling, medical and biological
sciences, engineering, and others. In the last few years, Graph Neural Networks (GNNs) have emerged as
a promising new supervised learning framework capable of bringing the power of deep representation
learning to graph and relational data. This ever-growing body of research has shown that GNNs achieve
state-of-the-art performance for problems such as link prediction, fraud detection, target-ligand binding
activity prediction, knowledge-graph completion, and product recommendations.
Deep Graph Library (DGL) is an open source development framework for writing and
training GNN-based models. It is designed to simplify the development of such models by using graph-
based abstractions while at the same time achieving high computational efficiency and scalability by
relying on optimized sparse matrix operations and existing highly optimized standard deep learning
frameworks (e.g., MXNet, PyTorch, and TensorFlow). This talk provides an overview of GNNs and DGL,
describes some recent developments related to high-performance multi-GPU, multi-core, and distributed
training, and describes our future development roadmap.
George Karypis is a Distinguished McKnight University Professor and an ADC Chair of Digital Technology at
the Department of Computer Science & Engineering at the University of Minnesota, Twin Cities. His
research interests span the areas of data mining, high performance computing, information retrieval,
collaborative filtering, bioinformatics, cheminformatics, and scientific computing. His research has
resulted in the development of software libraries for serial and parallel graph partitioning (METIS and
ParMETIS), hypergraph partitioning (hMETIS), for parallel Cholesky factorization (PSPASES), for
collaborative filtering-based recommendation algorithms (SUGGEST), clustering high dimensional datasets
(CLUTO), finding frequent patterns in diverse datasets (PAFI), and for protein secondary structure
prediction (YASSPP). He has coauthored over 280 papers on these topics and two books (“Introduction to
Protein Structure Prediction: Methods and Algorithms” (Wiley, 2010) and “Introduction to Parallel
Computing” (Publ. Addison Wesley, 2003, 2 nd edition)). In addition, he is serving on the program
committees of many conferences and workshops on these topics, and on the editorial boards of the IEEE
Transactions on Knowledge and Data Engineering, ACM Transactions on Knowledge Discovery from Data,
Data Mining and Knowledge Discovery, Social Network Analysis and Data Mining Journal, International
Journal of Data Mining and Bioinformatics, the journal on Current Proteomics, Advances in Bioinformatics,
and Biomedicine and Biotechnology. He is a Fellow of the IEEE.
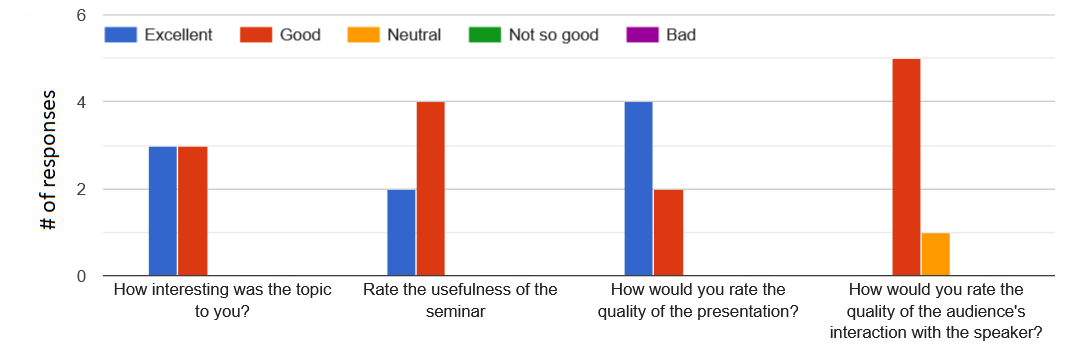
Selective Comments
"I found this talk really interesting and very pertinent to what I am currently working on."